Research interests
The dynamic conformation landscape of proteins often plays an important role in biological activity. Often, alterations to protein dynamics can lead to disruption of cellular homeostasis. During my PhD, I worked on the ubiquitination cascade, a post-translational modification mechanism in which a highly conserved protein, ubiquitin, is conjugated and deconjugated from substrate proteins, thereby altering their stability, localization, or activity. My work was focused on understanding the role of conformational dynamics of ubiquitin in its ability to get conjugated and deconjugated from substrates.
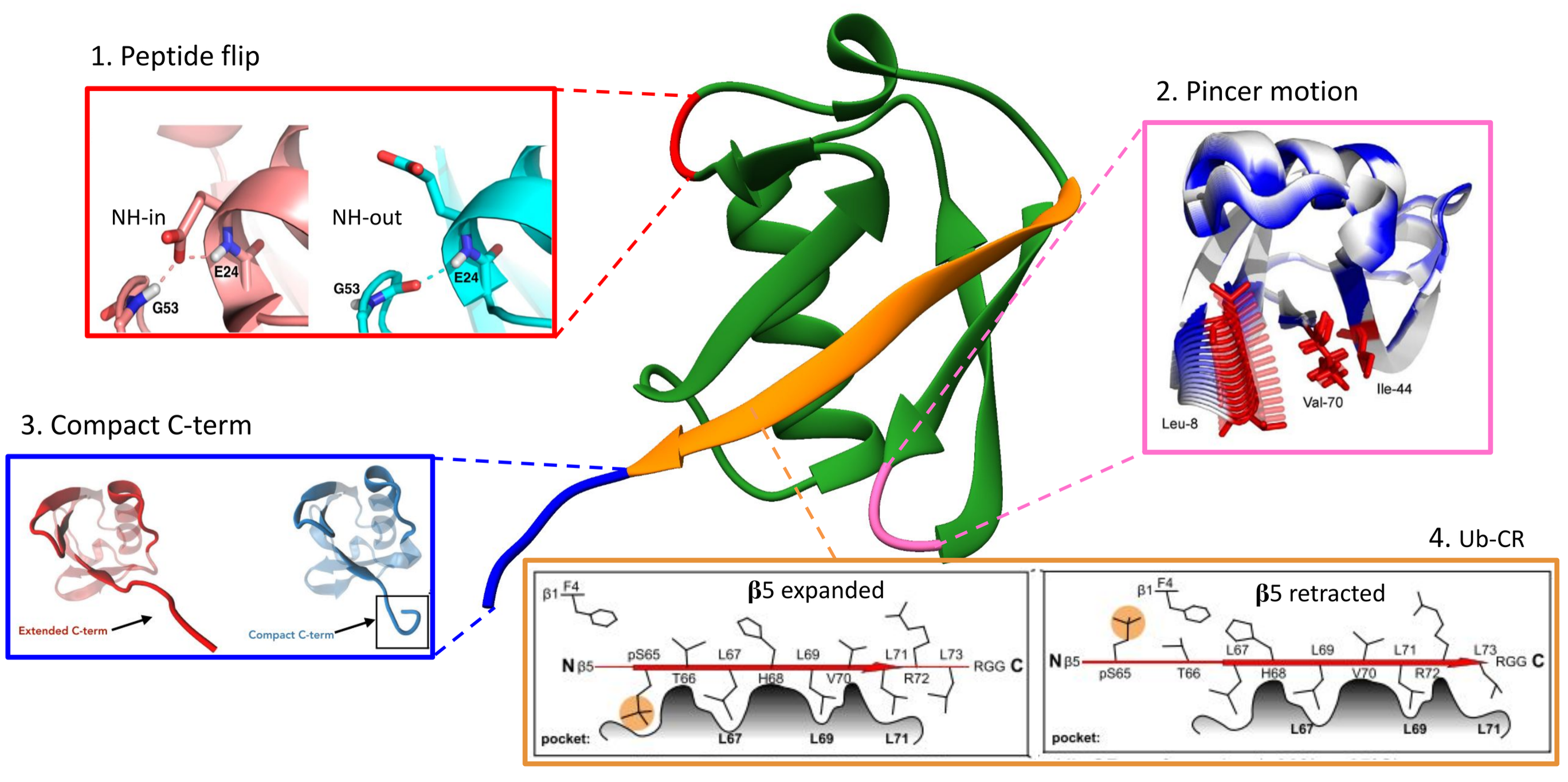
Since ubiquitin is at the center of a hub of reactions, various pathogens either hamper or co-opt the ubiquitination system to create permissive conditions for infection and replication. Baculoviruses encode a ubiquitin-like protein (viral ubiquitin), which retains the ubiquitin-like fold and conserved interaction interface. The deletion of viral ubiquitin hampers the maturation and assembly of the virion. Given the abundance of host ubiquitin, the function of viral ubiquitin was intriguing. This study showed that viral ubiquitin presents an altered conformational landscape, allowing it to differentially regulate the host ubiquitin machinery for successful infection.
Publications
Analyzing sub-millisecond timescale protein dynamics using eCPMG experiments
Journal of Biomolecular NMR, 2025
NMR methodology using extreme Carr-Purcell-Meiboom-Gill (eCPMG) experiments to characterize protein dynamics at sub-millisecond timescales. The technique allows investigation of rapid conformational changes in proteins.
The work includes:
- Development and application of eCPMG relaxation dispersion experiments
- Detection of fast protein dynamics at sub-millisecond timescales
- Methodology for studying conformational exchange processes
- Applications to structure-function relationships in biomolecules
The method provides tools to probe rapid protein dynamics and understand how proteins function through conformational changes.
Structural analysis of genetic variants of the human tumor suppressor PALB2 coiled-coil domain
Bioscience Reports, 2025
Here we solved the solution structure of the human PALB2 coiled-coil domain using NMR spectroscopy, which form an antiparallel homodimer. We then investigated how missense mutations affect the protein’s fold, homodimerization, and heterodimerization with BRCA1 to provide structural insights into reduced homologous recombination.
Key findings:
- Solution structure of PALB2 coiled-coil domain reveals an antiparallel homodimer
- Analysis of clinically relevant missense mutations and their structural implications
- Correlation between structural disruption and homologous recombination efficiency
- Approach for evaluating variants of uncertain significance in genetic counseling
